Hi everyone,
I’m back for a 3rd week of #PhDaily!
Considering the failure of last weeks ITC experiment, this week (and in the short term at least), I have decided to focus on my NMR experiments, as (fingers crossed) these are still working. I got into work at around 10 am, and set up my computer (a linux system), before being joined by my guide and NMR wizard Andrea.
Previously, I have looked at the backbone structure of the protein i am looking at (essentially a chain of carbon and nitrogen atoms). Now i am looking at the two other parts of the puzzle needed to get a protein structure, the side chains and the spacial analysis. Amino acids – the building blocks of proteins are varied, and some have different side chains which can affect the overall structure of a protein. The spacial analysis allows me to determine where the different amino acids are relative to each other in 3D space, which is crucial to produce a 3D structure of a protein.
There are two stages to each part of the NMR process, running the NMR scans, then processing the data. I am currently processing the side chain data, which like all NMR data, is like a giant confusing jigsaw puzzle!
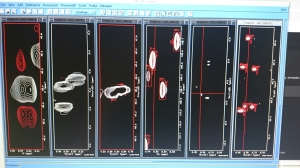
The NMR data processing screen. Certain peaks (the white circles) have to match up with certain other ones in each of these windows. From that i can tell the amino acid sequence, where all the side chains are, and where each amino acid is in relation to its neighbours.
I worked until 1pm, when i went for lunch and a sit outside, then back at 2pm for more data processing! I get my head around some difficult concepts to do with all of the different types of scan we have done, just in time to go home at 5.
Thanks for reading,
Microbe Stew

One thought on “#PhDaily Day 11 24/08/2015”